New BioGEOTRACES data sets: Connecting pieces of the microbial biogeochemical puzzle
Microorganisms play a central role in the transfer of matter and energy in the marine food web. Microbes depend on micronutrients (e.g. iron, cobalt, zinc, and a host of other trace metals) to catalyze key biogeochemical reactions, and their metabolisms, in turn, directly affect the cycling, speciation, and bioavailability of these compounds. One might therefore expect that marine microbial community structure and the functions encoded within their genomes might be related to trace metal availability in the ocean. The overall productivity of marine ecosystems—i.e. the amount of carbon fixed through photosynthesis—could in turn be influenced by trace metal concentrations.
For over a decade, the international GEOTRACES programme has been mapping the distribution and speciation of trace metals across vast ocean regions. Given the important relationship between trace metals and the function of marine ecosystems, biological oceanographers collaborate with GEOTRACES scientists to simultaneously probe the biotic communities at some sampling sites, allowing these biological data to be interpreted in the context of detailed chemical and physical measurements.
Two recent papers published in Scientific Data (see references below) describes two new, large-scale biological data sets that will facilitate studies aimed at understanding how microbes and metals relate to one another. Collected on four different sets of GEOTRACES cruises (see figure below), these papers report the public availability of hundreds of single cell genomes and microbial community metagenomes from the Pacific and Atlantic Oceans. The single cell genomes focus on the marine photosynthetic bacteria Prochlorococcus and Synechococcus and how they and other community members vary in different regions of the ocean. The metagenomic sequences provide snapshots of the entire microbial community found in each of these samples, yielding a broad overview of which microbes—and which genes, including those important for understanding nutrient cycling—are found in each sample. These two datasets are complementary and further enhanced by the wealth of chemical and physical data collected by GEOTRACES scientists on the same water samples. In particular, iron is of key interest, since it often limits primary productivity. These data sets can directly link iron availability with microbial community structure and gene content across ocean basins.
With these data, researchers can now ask questions such as how microbes have evolved in response to the availability or limitation of key nutrients and explore which organisms may be contributing to biogeochemical cycles in different parts of the global ocean. The extensive suite of chemical and physical measurements associated with these sequence data underscore their potential to reveal important relationships between trace metals and the microbial communities that drive biogeochemical cycles. These data sets also encourage cross-disciplinary collaborations and provide baseline information as society faces the challenges and uncertainties of a changing climate.
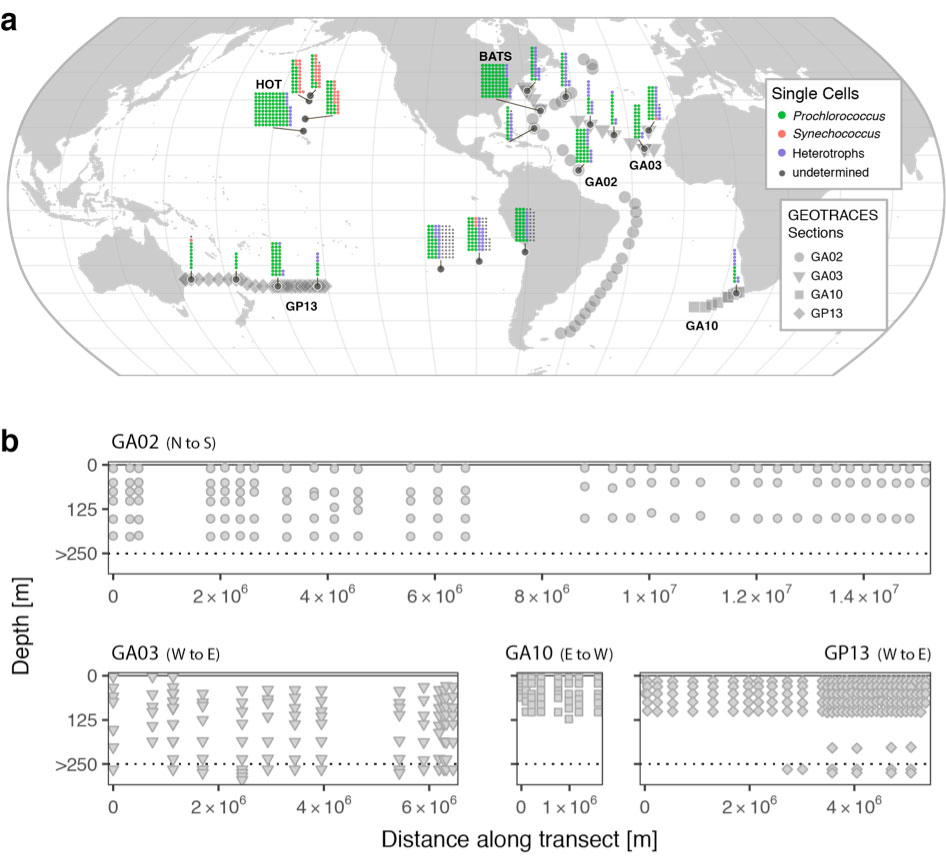
Figure: Locations and depths of samples. (a) Global map of sample locations. Single cell genomes are represented by miniaturized stacked dot-plots (each dot represents one single cell genome), with organism group indicated by color, and cells categorized as “undetermined” if robust placement within known phylogenetic groups failed due to low assembly completeness/quality or missing close references. Larger points correspond to stations on associated GEOTRACES sections where metagenomes were also collected. (b) Depth distribution of metagenome samples along each of the four GEOTRACES sections. Transect distances are calculated relative to the first station sampled in the indicated orientation. For clarity, the depth distribution of samples collected below 250 m are not shown to scale (ranging from 281–5601 m). Adapted from Berube et al. (2018) Sci. Data 5:180154 and Biller et al. (2018) Sci. Data 5:180176. Click here to view the figure larger.
Authors: Paul M. Berube (Massachusetts Institute of Technology), Steven J. Biller (Massachusetts Institute of Technology; current affiliation: Wellesley College) and Sallie W. Chisholm (Massachusetts Institute of Technology).
Published on Ocean Carbon & Biogeochemistry (OCB) December 2018 Newsletter.
References:
Berube, P. M. et al. (2018). Single cell genomes of Prochlorococcus, Synechococcus, and sympatric microbes from diverse marine environments. Scientific Data, 5, 180154. http://doi.org/10.1038/sdata.2018.154
Biller, S. J.,et al. (2018). Marine microbial metagenomes sampled across space and time. Scientific Data, 5, 180176. http://doi.org/10.1038/sdata.2018.176
